An extensive tutorial on electrostatic interactions for computer beginners, including Python exercises.
On this page
What are Electrostatic Interactions?
Imagine you’ve just taken a sweater out of the dryer and it’s sticking to your shirt. That “static cling” is a perfect example of an electrostatic interaction.
In the world of proteins, atoms can have electrical charges. Just like magnets:
- Opposite charges attract: A positive (+) atom and a negative (-) atom will pull toward each other.
- Like charges repel: Two positive (+) atoms or two negative (-) atoms will push each other away.
In proteins, these interactions are often called Salt Bridges when they happen between specific “charged” amino acids (like Lysine and Aspartate). They act like tiny internal staples that keep the protein’s shape stable.
Electrostatic interactions are much stronger than Van der Waals forces but can be weakened if the protein is surrounded by water or salt.
Introduction
Two atoms that are electrically charged interact electrostatically with each other.
The nature of these interactions depends on the signs of the charges; charges of the same sign repel each other, and charges of opposite sign attract each other.
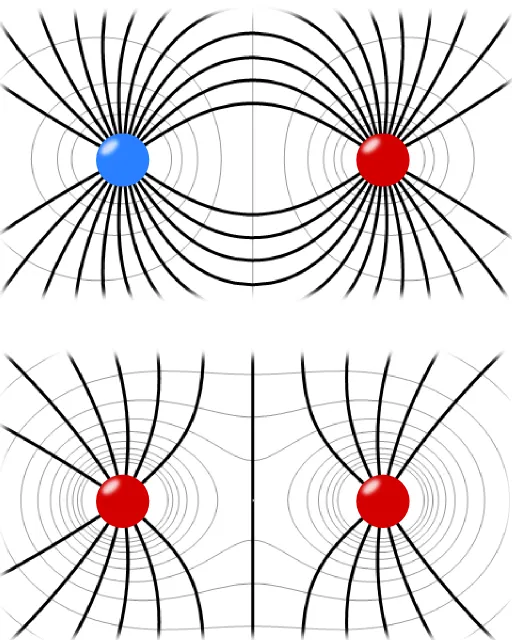
Each of the interacting charges generates an electric field, which surrounds it.
The interaction between the two charges results from the way the electric field emanating from one charge affects the other charge.
Since both full and partial charges may be involved, there are various types of electrostatic interactions between atoms, including charge-charge, charge-dipole, and dipole-dipole, where the latter two may involve fixed or induced dipoles.
In proteins, the most common interactions (except for Interaction - van der Waals interactions) are:
-
Ionic interaction occurs between fully charged atoms. When the atom-atom distance is less than 4 Å, the ionic interaction is referred to as an ‘ion pair’ or ‘salt bridge’.
-
Hydrogen ‘bond’ occurs between two electric dipoles, one of which includes a hydrogen atom. Less standard hydrogen bonds may involve a full charge and a dipole.
Electrostatic interactions, and hydrogen bonds in particular, are very common in proteins, and play different roles, such as imparting specificity to the protein structure or to its association with other molecules; contributing to protein folding; and aiding in enzymatic catalysis.
In proteins, these interactions may occur as follows:
- Between charged atoms belonging to different chemical groups within the protein.
- Between charged atoms on the protein surface and charges in the environment (e.g., the cytoplasm, membrane, interstitial fluid, or other bodily fluids).
- Between charged atoms of the protein and those present on its ligand.
Coulomb interactions
The potential energy of interaction between two point charges in a uniform medium is described by Coulomb’s law:
(where is the potential interaction energy (multiply by 332 (kcal/mol) to obtain the energy in kcal/mol), and are the interacting charges (in electron charges), is the distance between the charges (in Å), and is the relative dielectric constant of the medium).
The potential energy of a noncovalent interaction describes the energy resulting from interaction between two species (atoms, groups, molecules) in a static system. It is therefore only one component of the total (i.e., free) energy of the system; the latter accounts also for changes in the interaction resulting from dynamics of both the interacting species and their environment (i.e., entropic changes).
Equation demonstrates the three quantities on which the interaction energy depends:
- : The magnitude of the interacting charges.
- : The distance between the charges.
- : The dielectrics of the medium.
The dependency of the interaction on the second and third quantities provides important information on its physical nature.
Note that although and may change the strength of the Coulomb interaction, they cannot change its nature, i.e., attractive or repulsive. That depends only on the signs of the interacting charges.
Distance
The energy depends inversely on the distance between the charges ().
That is, the charges must be separated by a relatively large distance in order for the interaction to become insignificant.
Indeed, electrostatic interactions in proteins may be significant even at distances of up to a few tens of Ångströms. By ‘significant’ we mean an energy value equal to or greater than (≈ 0.6 kcal/mol).
Medium
The dielectric constant () represents the ability of the medium to “mask” (i.e., reduce) the energy of the interaction between charges.
Analogy: Imagine two people trying to talk to each other.
- In a vacuum (), it’s like a quiet room—they can hear each other perfectly. The interaction is maximal.
- In water (), it’s like a loud, crowded party. The noise (the high dielectric constant) makes it very hard for them to communicate. The interaction is 80 times weaker!
The dependency of electrostatic interactions on the dielectric constant provides a great deal of information about the environmental aspect of the interactions.
The dielectric constant represents the ability of the medium to mask (i.e., reduce) the energy of the interaction between charges:
-
In a vacuum, there is no masking at all, and the dielectric constant is 1.
-
Water represents the other extreme; there is ample masking of electrostatic interactions, as reflected by the high dielectric constant (~80). The electrostatic interactions in water will therefore be 80 times weaker than in a vacuum.
-
The cytoplasm of cells is generally considered to be an aqueous, high-dielectric environment. However, measurements indicate that the dielectric constant inside cells is ~60 at the most.
-
Lipid media, such as biological membranes, have dielectric constants of 2 to 30, higher than that of a vacuum but still considered low.
Thus, the Coulomb interaction in cytoplasm and lipid media will be considerably stronger than in water. Even inside the protein, the dielectrics may change, which makes things much more complicated.
Python Exercise: Calculating Electrostatic Energy
In this exercise, we will use Python to calculate the actual interaction energy between two charged atoms in a protein structure. We will use the formula:
1. Setup
If you haven’t already, install ProDy and numpy:
uv init electrostatic-projectcd electrostatic-projectuv add prody numpy2. The Python Script
Create a file named calc_electrostatic.py. We will calculate the interaction between a Lysine (positive) and an Aspartate (negative) in the protein Ubiquitin (1UBQ).
from prody import parsePDBimport numpy as np
# 1. Load the protein (Ubiquitin)pdb = parsePDB('1ubq')
# 2. Select two atoms that form a salt bridge# Atom NZ in Lysine 27 (Positive charge +1)# Atom OD1 in Aspartate 52 (Negative charge -1)lys_nz = pdb.select('resnum 27 and name NZ')asp_od = pdb.select('resnum 52 and name OD1')
if lys_nz and asp_od: # 3. Get coordinates pos1 = lys_nz.getCoords()[0] pos2 = asp_od.getCoords()[0]
# 4. Calculate distance (r) in Angstroms r = np.sqrt(np.sum((pos1 - pos2)**2)) print(f"Distance between Lys27(NZ) and Asp52(OD1): {r:.2f} Å")
# 5. Physics constants q1 = 1.0 # Lysine is +1 q2 = -1.0 # Aspartate is -1 k = 332 # Conversion factor to kcal/mol
# We will check energy in two different environments: dielectrics = {'Vacuum': 1, 'Protein Interior': 4, 'Water': 80}
print("\nCalculated Interaction Energy (kcal/mol):") for environment, epsilon in dielectrics.items(): # Coulomb's Law formula energy = k * (q1 * q2) / (epsilon * r) print(f"- In {environment} (ε={epsilon}): {energy:.2f} kcal/mol")
else: print("Could not find the specified atoms.")3. Understanding the Results
- Distance (): If the distance is under 4.0 Å, it’s considered a strong salt bridge.
- The Sign: Notice the energy is negative. In physics, negative energy means the atoms are attracted to each other (they are stable).
- The Environment: Look at how the energy drops from -80 kcal/mol (very strong) in a vacuum to only -1 kcal/mol in water. This shows why salt bridges are much stronger inside a protein where water is excluded!
Challenge for You
- Find another pair: Look for
resnum 11(Lysine) andresnum 3(Aspartate). Are they close enough to interact? - Change the charges: What happens if you change
q2to1.0(simulating two positive charges)? Does the energy become positive (repulsive)? - Graph it: If you know how to use
matplotlib, try plotting the energy vs. distance from 2.0 Å to 10.0 Å.
Further Reading
- Interaction - Hydrogen bond - A special type of electrostatic interaction.
- Interaction - van der Waals - The much weaker “neutral” attraction.